Functions and structural dynamics of nuclear pore complexes (NPC)
Real time visualization of the dynamics of nano-machine gatekeepers at the surfaces of cell membranes

Richard Wong, Professor, Nano Life Sciences Institute (WPI-NanoLSI), Kanazawa University
Nanomachines as gatekeepers of molecular transport across cell membranes
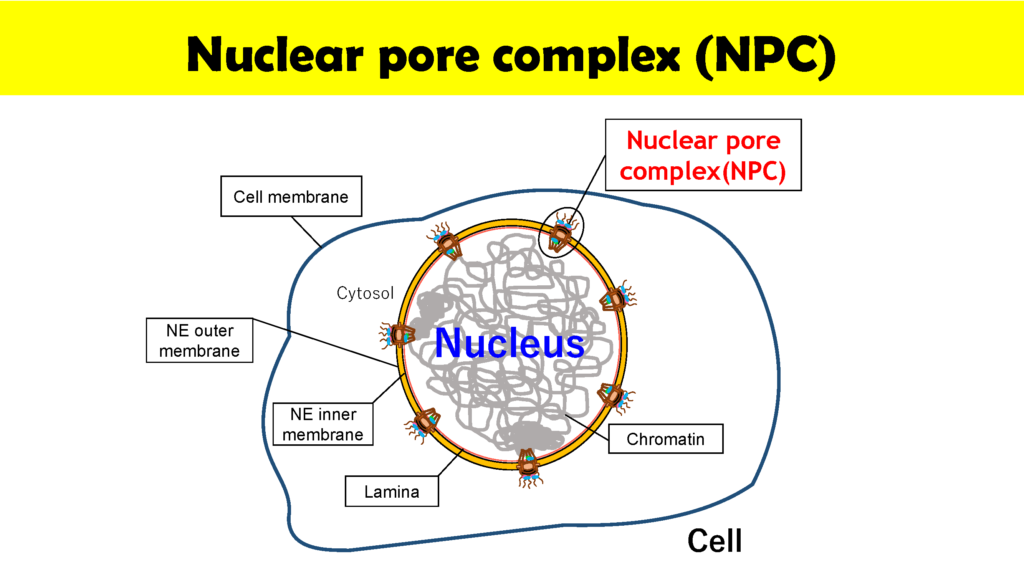
Nuclear pore complexes (NPC) at the surfaces of nuclear membranes play a critical role in regulating the transport of molecules between the cell nucleus and cytoplasm. “NPCs can be thought of as nanomachines composed of about 30 proteins that we call nucleiporins,” says Wong. “The important point is that NPCs are gatekeepers that allow the movement of preferred molecules such as RNA and proteins across the cell membrane as well as preventing undesirable molecules that may cause disease and cell death, from making the crossing. But in spite of the importance of NPCs we still only have a limited understanding of the dynamics of NPCs. My goal is to utilize Kanazawa University’s internationally renowned high speed atomic force microscopy (HS-AFM) technology to visualize the dynamics of NPCs in real time.”
To-date the physical properties NPC have largely been studied by electron microscopy using static samples. In contrast, Wong and his group is collaborating with colleagues at the Kanazawa WPI-NanoLSI to directly image the functions and structural dynamics of NPCs. “The NPC is the last line of defense in cells to prevent infection and viral replication,” explains Wong. “We are focusing on imaging inner channels of NPC in cancer cells.”
Delving into the mysteries of NPCs with scanning nano-probes
Specific topics include studies on disruption and formation process of the nuclear pore complex, and formation of hemochromatin directly under the nuclear membrane and nuclear pore complex. The instrumentation being used includes ultra-high resolution optical microscope for observing ultra-fine structures; confocal microscope for observing cell structures; and high speed AFM technology for protein dynamics. With this goal, Wong and colleagues used HS-AFM to observe native NPCs. The measurements were made in the tapping mode of the HS-AFM with the cantilever moving with at 2 nm free amplitude at 900 kHz and 200 frames/s. The images of the nuclear envelope surface and single NPCs showed both plugged and free NPCs.
Wong has also used HS-AFM to visualize the dynamics of NPC in colon cancer cells [1]. Specifically, this “nanoendoscopic” approach was used to determine the behavior of FG (phenylalanine-glycine)-nucleoporins (FG-Nups) that constitute the central turnstile enclosed by the cytoplasmic filaments. Images showed the FG-Nups to be “short, hair-like, twisted ropes having a brush-like manner or forming a broken cobweb shape from their tethering point”.
“We describe this structure as being ‘spiders-spidermen working inside a sticky cobweb’,” says Wong. “This is our model for the process of central channel shuttling or gene gating by the nuclear pore.” These findings offer potential for the development of tailored drug delivery for targeting cancer cells.
Future challenges include synthesizing artificial 100 nm pores for modeling protein trafficking. “Put simply, we will chop up proteins and put them inside cells,” says Wong. “But the actual process is non-trivial, but is very important for looking inside NPCs.”
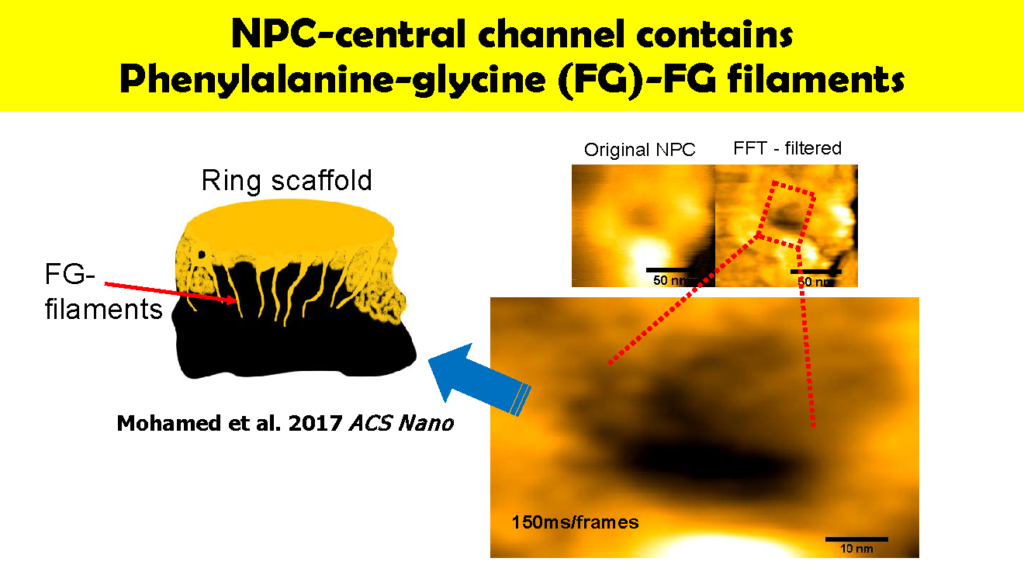

(left)Original NPC
(right)FFT – filtered

150ms/frames
Research highlights
Recent nanopore NPC related research papers include, “High-speed atomic force microscopy reveals loss of nuclear pore resilience as a dying code in colorectal cancer cells”, published in ACS Nano [1], “ROCK-dependent phosphorylation of NUP62 regulates p63 nuclear transport and squamous cell carcinoma proliferation”, published in EMBO reports [2], “Disease-specific alteration of karyopherin-α subtype establishes feed-forward oncogenic signaling in head and neck squamous cell carcinoma” published in Oncogene [3] and “Nucleoporin translocated promoter region (TPR) upregulation alters mTOR-HSF1 trails and suppresses autophagy induction in ependymoma” published in Autophagy.
References
- MS Mohamed et al, “High-Speed Atomic Force Microscopy Reveals Loss of Nuclear Pore Resilience as a Dying Code in Colorectal Cancer Cells”, ACS Nano. 11, pp 5567–5578 (2017). doi: 10.1021/acsnano.7b00906.
- M Hazawa et al, “ROCK‐dependent phosphorylation of NUP62 regulates p63 nuclear transport and squamous cell carcinoma proliferation”, EMBO Rep. 19(1):73-88 (2018). doi: 10.15252/embr.201744523.
- Hazawa et al, “Disease-specific alteration of karyopherin-α subtype establishes feed-forward oncogenic signaling in head and neck squamous cell carcinoma”, Oncogene (2019). doi: 10.1038/s41388-019-1137-3. [Epub ahead of print]
- Dewi et al, “Nucleoporin translocated promoter region (TPR) upregulation alters mTOR-HSF1 trails and suppresses autophagy induction in ependymoma.”, Autophagy (2020) accepted

