Selection ‘Hit’-to-Clinical ‘Fit’: Potential of Peptamer for drug discovery
Innovative screening, verification, and optimization technology for analyzing millions of aptamers in the search for next generation cancer therapeutics

Madhu Biyani, Assistant Professor
Drug discovery has a long history. Notable, milestones include the development and proliferation of small-molecule drugs towards the end of the 19th century but these drugs were found to induce side effects leading to the search for new ideas and approaches. The next phase of development led to the advent of so-called ‘biologics-drugs’ as exemplified by antibodies produced from living organisms. These drugs were safe and exhibited high specificity but the number of targets in the body—so called biomarkers—far outnumbered the number of known antibodies. Next came peptides—short proteins consisting of 2 to 50 amino acids—found in eggs, milk, and meat, and other such sources. Now, in the 21st century, scientists have turned their attention to exploiting the unique biochemical properties of ‘aptamers’ by synthesizing artificial, 3D structures of short peptides referred to as ‘peptamers’ that exhibit the properties of finding, binding and inhibiting specific target molecules with remarkable sensitivity. Notably, aptamers are considered to be synthetic alternatives to natural antibodies found in the human body and are sometimes referred to as ‘chemical antibodies’.
“In its simplest form, drug discovery involves target identification and validation, searching for aptamer molecules that bind to the target, and finally clinical trials to test whether the aptamer molecules bind and nullify the action of target molecules, for example, to prevent harmful tumors,” explains Madhu Biyani, an assistant professor at the Nano Life Science Institute (WPI-NanoLSI) who joined Kanazawa in the Spring of 2020. “The most time consuming step in this process is identifying potentially effective ‘hit’ aptamer molecules from millions of potential candidates to use in clinical trials. This process of aptamer screening and evaluation is my core area of research. We often refer to the three phases as screening (Hit), verification (Fit), and Optimization (Sit). It’s a very exciting and dynamic area of interdisciplinary research. I am looking forward to using my experience in aptamer screening to join the world renowned AFM-team at nanoLSI to work towards achieving the goals of the WPI at Kanazawa University.”
Targeting cancer biomarkers
- Protease Cathepsin E (CTSE), a prognostic marker in cancer
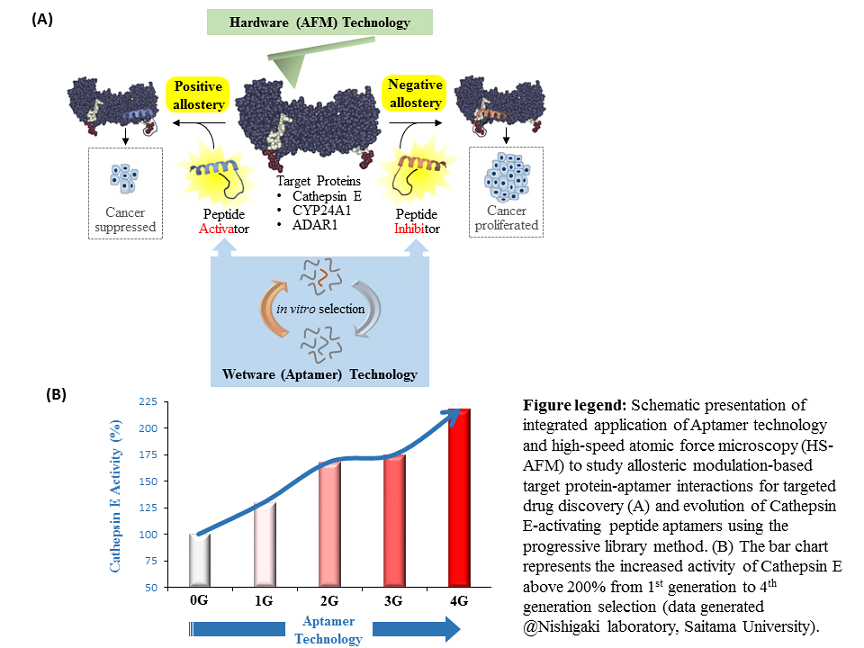
Cathepsin E (CTSE) is a protease that acts as a catalyst to breaks down proteins into amino acids. Importantly, CTSE has emerged as a therapy target for cancers and shows potential for early cancer diagnosis based on experimental research showing its active role in maintaining homeostasis and immune response of the human body. “In one of earlier JST/MEXT project at Saitama University, headed by Prof Nishigaki and team including me, identified novel candidates of peptamer for CTSE,’’ says Biyani. The team have identified ‘Hit’ peptamers using their advanced ‘in-vitro selection technology’ yielding CTSE-activity enhancement of over than 200%. Now, I look forward to take these ‘Hit’ peptamers in advancing to the phases ‘Fit’ and ‘Sit’ with the experts in AFM technology at the NanoLSI,” says Biyani.
- Enzyme CYP24A1 and cancer therapy
Research has shown that over expression of the enzyme CYP24A1 is linked to the start of cancer and other ailments such as chronic kidney disease and osteoporosis. In collaboration with Prof Yasuda at Toyama Prefectural University, Biyani is using her in-vitro screening technology to develop CYP24A1 aptamer inhibitors to enhance the physiological functions of calcitriol for cancer treatment and prevention. “Our approach is different from the conventional SELEX method and we integrated this with DEPSOR, a portable and rapid electrochemical sensing platform,” says Biyani.
- Action of adenosine deaminases on RNA1 (ADAR1)
Overexpression of the enzyme ADAR1 leads to the promotion of lung, liver and, other forms of cancer, so scientists are developing strategies to target it as a form of cancer therapy. Biyani and her colleagues are developing a new fast and sensitive in-vitro electrochemical RNA editing assay using DEPSOR to identify potential aptamer candidates that inhibit ADAR1 deaminase activity. “This theme of ‘elucidation of dynamic structural property of ADAR1 in a process of A-to-I RNA editing for development of ADAR1 inhibitors for cancer therapy’ was awarded a Transdisciplinary Research Promotion Grant by the NanoLSI this year,” says Biyani. “I am looking forward to developing the ADAR1-inhibiting aptamer molecules in collaboration with NanoLSi colleagues including Professors Nakajima and Kodera.”
Women in science
Assistant Professor Biyani is from India and has been in Japan since 2000. Originally trained as a physician, she changed to a research career after coming to Japan and in 2011 completed a doctorate in bioengineering at Saitama University. “Now, I am very fortunate to be able to have the opportunity to work under the guidance of Professor Miki Nakajima who is a PI at NanoLSI and received the ‘2019 Haazami Female Researcher Award’ for her contributions to drug metabolism and drug discovery. Japan is a second home for me. And the NanoLSI is a stimulating environment for research. The PIs and administration staff are very helpful, and I am grateful for their wide ranging of support. I am really looking forward to my pursuing my research at the NanoLSI at Kanazawa University.”
Selection of publications
- Madhu Biyani et al., Instant enumeration of total viable bacterial counts for food quality assurance using ‘DEP-On-Go’ sensor, Analytical Methods 10, 1585, (2018). https://doi.org/10.1039/c7ay02927f
- Manish Biyani et al., PEP-on-DEP: A competitive peptide-based disposable electrochemical aptasensor for renin diagnostics, Biosensors and Bioelectronics 84, 120-125, (2016). DOI: doi.org/10.1016/j.bios.2015.12.078