Watching Proteins Move with SimHS-AFMfit-MD: A New Method Reconstructs Dynamic Atomic Structures of Proteins
Overview
Proteins are not rigid molecular sculptures. They are soft mechanical systems.
They bend, breathe, hinge, and twist as they carry out life’s chemistry — regulating signaling, assembling cellular structures, and generating force. Yet for decades, structural biology has faced a fundamental limitation: most experimental techniques reveal structure, but not motion or dynamics.
X-ray crystallography and cryo-electron microscopy provide exquisite atomic detail, but only as static snapshots — molecules frozen in time. In contrast, molecular dynamics (MD) simulations can model continuous motion at atomic resolution, yet simulations alone cannot guarantee that the predicted motions actually occur in real biological conditions.
In this study, an interdisciplinary collaboration of biochemists, biophysicists, and computational scientists led by Dr. Kien Xuan Ngo and Dr. Takashi Sumikama (Nano Life Science Institute, Kanazawa University, Japan; IFIRSE, Quy Nhon, Vietnam), together with Dr. Rémi Vuillemot and Dr. Sergei Grudinin (Univ. Grenoble Alpes, France), introduces a new solution. By combining high-speed atomic force microscopy, flexible structural modeling (AFMfit), and all-atom MD simulations, the team developed SimHS-AFMfit-MD, an integrative framework that converts live nanoscale imaging into atomic-resolution molecular dynamics.
Rather than predicting protein motion or merely imaging it, the method infers atomic motions directly from experimental movies.
Using the actin crosslinking protein alpha-actinin as a model system, the researchers transformed thousands of high-speed AFM images into ensembles of atomic conformations. The reconstructed dynamics revealed structural transitions previously inaccessible to direct observation, including distinct conformational behaviors between calcium-bound and calcium-free states.
The work closes a longstanding spatial resolution gap between experimental observation and physical simulation, allowing molecular motion to be visualized while it is occurring.
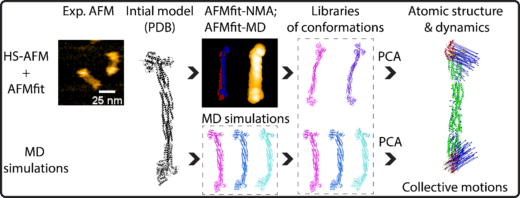
Figure A novel orkflow for analyzing the dynamics and atomic structures of alpha-actinin through the integration of HS-AFM with the AFMfit-NMA, AFMfit-MD, and MD simulations. The produced 3D conformations were processed by principal component analysis (PCA) to identify and classify dynamic atomic structures into distinct modes of collective motion (indicated by blue arrows). Copywrite: © 2026 Ngo et, published by American Chemical Society (CC-BY-NC-ND 4.0 ).
What the Researchers Achieved
The SimHS-AFMfit-MD framework operates by merging complementary sources of information, each incomplete on its own but powerful together.
High-speed AFM provides real-time movies of single protein molecules under near-physiological conditions, capturing their surface conformations during they fluctuate. However, AFM measures only topography — not internal atomic structure.
To interpret these images, the team applied AFMfit flexible modeling using nonlinear normal mode analysis (NMA), which deforms atomic models in physically plausible ways to match observed shapes. For the first time in this field, molecular dynamics simulations were then used to guide and validate the fitting, constraining the solutions to physically realistic motions.
This pipeline reconstructs three-dimensional atomic conformations from surface images — a task long considered impossible because surface measurements lack direct atomic information.
Cross-validation against unbiased all-atom MD simulations confirmed that the inferred motions accurately represent real molecular behavior. The analysis revealed coordinated bending and torsional motions of alpha-actinin and showed how calcium binding stabilizes specific structural states, a key regulatory mechanism in cytoskeletal organization.
Why It Matters
The central challenge in contemporary structural biology is no longer simply determining molecular structures — it is understanding dynamic function.
Many biologically important systems, including intrinsically disordered proteins, signaling complexes, and cytoskeletal assemblies, do not exist as single structures but as continuously shifting ensembles. Static methods struggle to describe such molecules because function often emerges precisely from their motion.
SimHS-AFMfit-MD addresses this problem by:
- transforming live microscopy movies into atomic structural ensembles
- experimentally constraining molecular simulations
- enabling direct observation of conformational transitions in solution
In effect, the approach establishes a new experimental category: dynamic atomic structural biology — the ability to observe how proteins operate rather than merely what their surfaces look like.
The results also provide a dynamic interpretation of previously solved crystallographic and cryo-EM structures, explaining how those static states are connected during real biological activity.
Future Directions
The implications extend beyond a single protein system.
The conformational libraries generated by SimHS-AFMfit-MD can serve as experimental benchmarks for improving molecular simulation force fields and as training datasets for machine-learning models that aim to predict protein dynamics across timescales, thereby bridging the temporal resolution gaps between experimental imaging and simulation methods. Integration with emerging high-speed 3D-AFM imaging may allow dynamic reconstruction of large molecular machines and flexible multi-protein assemblies.
More broadly, the framework demonstrates a new paradigm: microscopy and simulation are no longer competing approaches but complementary components of a single measurement.
By unifying real-time imaging with physics-based modeling, the method moves structural biology toward a long-standing goal — not simply determining protein structure, but understanding protein behavior in real time at atomic resolution.
Article
- Title
- SimHS-AFMfit-MD: An Integrative Approach for Inferring Alpha-Actinin Atomic Conformational Dynamics
- Author
- Kien Xuan Ngo*, Takashi Sumikama*, Rémi Vuillemot*, Han Gia Nguyen, Ngan Thi Phuong Le, Sergei Grudinin* (*Corresponding authors)
- Journal
- Nano Letters
- Publication date
- Feb 25, 2026
- DOI
- 10.1021/acs.nanolett.6c00617
- URL
- https://pubs.acs.org/doi/10.1021/acs.nanolett.6c00617

